 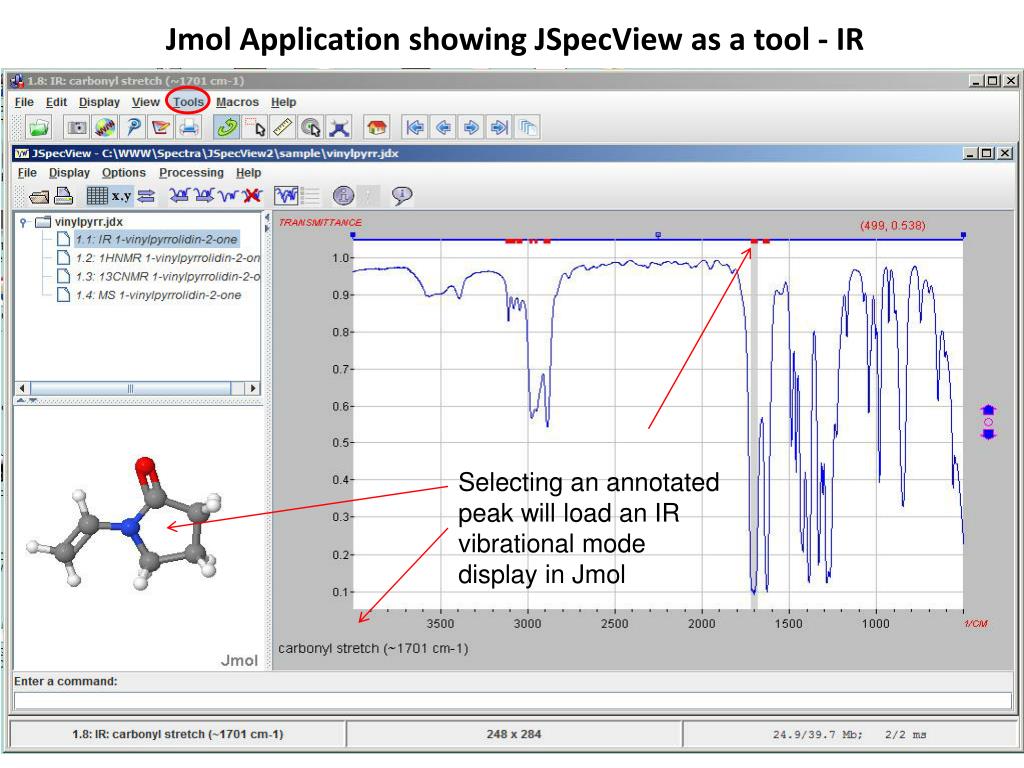
Jmol/JSmol is ideal for development of Web-based courseware and Web-accessible chemical databases. Jmol interfaces well with JSpecView for spectroscopy, JSME for 2D->3D conversion, POV-Ray for images, and CAD programs for 3D printing (VRML export). The default viewer can be configured in the Structure tab in the ToolsPreferences dialog box. ChimeraX and PyMOL support is included from Jalview 2.11.2. Jalview 2.8.2 included support for Chimera, provided it is installed and can be launched by Jalview. Advance to the last frame because the quantum software puts the optimized orbital information in the last frame. Features include interactive animation and linear morphing. The Jmol viewer has been included since Jalview 2.3. A rich scripting language and a well-developed Web API allow easy customization of the user interface. sequence id pop-up menu associate a PDB structure internal pdb viewer. Multiple files can be loaded and compared. Features include reading a variety of file types and output from quantum. The J in Jmol stands for Java, but we want to use it in a web. Files can be transferred directly from several databases, including RCSB, EDS, NCI, PubChem, and MaterialsProject. Jmol is a Java molecular viewer for three-dimensional chemical structures. When you need a molecule viewer, you should use Jmol unless you have a very good reason not to. Jmol can read many file types, including PDB, CIF, SDF, MOL, PyMOL PSE files, and Spartan files, as well as output from Gaussian, GAMESS, MOPAC, VASP, CRYSTAL, CASTEP, QuantumEspresso, VMD, and many other quantum chemistry programs. Jmol/JSmol is a molecular viewer for 3D chemical structures that runs in four independent modes: an HTML5-only Web application utilizing jQuery, a Java applet, a stand-alone Java program (Jmol.jar), and a "headless" server-side component (JmolData.jar). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
